fitting example: fitting_t1_data¶
This example shows how to use nmrglue and scipy optimize module to fit T1 relaxation trajectories. Three scripts are used in the process.
First the extract_trajs.py script reads in box limits from boxes.in and a list of spectra from spectra.in. The script integrates each peak in each spectrum and writies the trajectory for each peak to disk as traj.npy in numpy .npy format.
#! /usr/bin/env python
# Scipt to extract trajectories from a series a 2D spectrum.
import nmrglue as ng
import numpy as np
# read in the integration limits and list of spectra
peak_list = np.recfromtxt("boxes.in")
spectra_list = np.recfromtxt("spectra.in")
# prepare the trajs records array
num_spec = spectra_list.size
num_peaks = peak_list.size
elist = [np.empty(num_spec,dtype="float") for i in xrange(num_peaks)]
trajs = np.rec.array(elist,names=list(peak_list.f0))
# loop over the spectra
for sn,spectra in enumerate(spectra_list):
# read in the data from a NMRPipe file
print "Extracting from:",spectra
dic,data = ng.pipe.read(spectra)
# loop over the integration limits
for name,x0,y0,x1,y1 in peak_list:
# integrate the region and save in trajs record array
if x0>x1: x0,x1 = x1,x0
if y0>y1: y0,y1 = y1,y0
trajs[name][sn] = data[y0:y1+1,x0:x1+1].sum()
# normalize each trajectory
for peak in trajs.dtype.names:
trajs[peak] = trajs[peak]/trajs[peak].max()
# save the trajectories records array to disk
np.save("traj.npy",trajs)
[boxes.in]
#Peak X0 Y0 X0 Y1
A20 4068 938 4079 913
A24 3992 1013 4000 997
A26 4065 962 4075 940
A34 4009 985 4018 958
A48 4028 1034 4036 1010
C28 4035 1115 4044 1092
D36 3994 987 4003 973
D40 4076 802 4085 774
D46 4155 899 4163 883
D47 4053 967 4062 941
E15 4162 1022 4170 996
E19 4176 902 4185 875
E27 4036 1084 4044 1054
E42 4136 1055 4142 1026
E56 4107 821 4115 794
F30 4013 1060 4023 1031
F52 4097 828 4105 799
G09 4054 1249 4063 1220
G14 4068 1331 4077 1304
G38 4098 1254 4106 1227
G41 4091 1283 4099 1259
I06 4087 903 4096 884
data/Ytau_100.fid/test.ft2
data/Ytau_100000.fid/test.ft2
data/Ytau_250000.fid/test.ft2
data/Ytau_500000.fid/test.ft2
data/Ytau_750000.fid/test.ft2
data/Ytau_1000000.fid/test.ft2
data/Ytau_1500000.fid/test.ft2
data/Ytau_2000000.fid/test.ft2
data/Ytau_3000000.fid/test.ft2
data/Ytau_4000000.fid/test.ft2
The second script fit_exp_leastsq.py reads in this traj.npy file and the T1 relaxation times associated with the spectra collected from time.dat. Each trajectory is fit using the least squares approach. Other optimization algorithms can be substituted with small changes to the code, see the scipy.optimize <http://docs.scipy.org/doc/scipy/reference/optimize.html>_ documentation). The resulting fits are saved to a fits.pickle file for easy reading into python as well as the human readable fits.txt file.
#! /usr/bin/env python
# fit a collection to T1 trajectories to a decaying exponential
import scipy.optimize
import numpy as np
import pickle
# read in the trajectories and times
trajs = np.load("traj.npy")
t1 = np.recfromtxt("time.dat")
# fitting function and residual calculation
def fit_func(p,x):
A,R2 = p
# bound A between 0.98 and 1.02 (although fits do not reflect this)
if A>1.02:
A = 1.02
if A<0.98:
A = 0.98
return A*np.exp(-1.0*np.array(x)*R2/1.0e6)
def residuals(p,y,x):
err = y-fit_func(p,x)
return err
p0 = [1.0,0.05] # initial guess
fits = {}
# loop over the peak trajectories
for peak in trajs.dtype.names:
print "Fitting Peak:",peak
# get the trajectory to fit
traj = trajs[peak]
# fit the trajectory using leastsq (fmin, etc can also be used)
results = scipy.optimize.leastsq(residuals,p0,args=(traj,t1))
fits[peak] = results
# pickle the fits
f = open("fits.pickle",'w')
pickle.dump(fits,f)
f.close()
# output the fits nicely to file
f = open("fits.txt",'w')
f.write("#Peak\tA\t\tR2\t\tier\n")
for k,v in fits.iteritems():
f.write(k+"\t"+str(v[0][0])+"\t"+str(v[0][1])+"\t"+str(v[1])+"\n")
f.close()
[time.dat]
# time in us
100
100000
250000
500000
750000
1000000
1500000
2000000
3000000
4000000
Results:
[fits.txt]
#Peak A R2 ier
E19 0.992583088163 0.165456904924 1
D36 0.978162369609 0.150387170038 1
E15 0.996022817946 0.0881391067001 1
G41 0.884746148818 0.528402076365 1
G09 0.993838313697 0.0974310406654 1
E56 0.978642097098 0.153694397368 1
D40 0.937320157503 0.32803199035 1
A34 1.00238376034 0.152055981221 1
E42 0.97955812576 0.201548076316 1
E27 0.957068222368 0.290667669811 1
F52 1.01437765462 0.063087299437 1
G38 1.00575140025 0.127826452523 1
C28 0.779226606115 0.680794315854 1
F30 0.955609064219 0.258560125202 1
G14 0.994898872827 0.108697405902 1
D47 0.993890875485 0.101193820532 1
D46 0.978622370865 0.074505374897 1
A48 0.984906624566 0.125792739083 1
A20 0.99998574293 0.102821608635 1
A24 0.923616582091 0.408644672104 1
A26 0.964157015536 0.252957335373 1
I06 0.997427797051 0.0604968616212 1
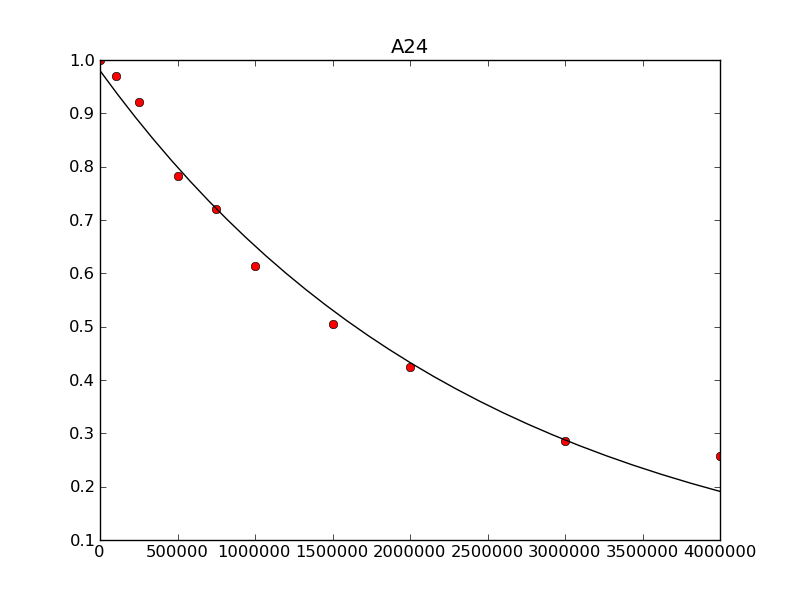
The last script pt.py reads in the fits, trajectories and T1 relaxation times and plots the experimental points and best fit to a series of *_plot.png files.
[pt.py]
#! /usr/bin/env python
# Plot trajectories and fitting results
import pickle
import matplotlib.pyplot as plt
import numpy as np
# the same fit_func as in fit_exp_leastsq.py
def fit_func(p,x):
A,R2 = p
# bound A between 0.98 and 1.02 (although fits do not reflect this)
if A>1.02:
A = 1.02
if A<0.98:
A = 0.98
return A*np.exp(-1.0*np.array(x)*R2/1.0e6)
# read in the trajectories, fitting results, and times
fits = pickle.load(open("fits.pickle"))
trajs = np.load("traj.npy")
times = np.recfromtxt("time.dat")
sim_times = np.linspace(times[0],times[-1],2000)
# loop over the peaks
for peak,params in fits.iteritems():
print "Plotting:",peak
exp_traj = trajs[peak]
sim_traj = fit_func(params[0],sim_times)
# create the figure
fig = plt.figure()
ax = fig.add_subplot(111)
ax.plot(times,exp_traj,'or')
ax.plot(sim_times,sim_traj,'-k')
ax.set_title(peak)
# save the figure
fig.savefig(peak+"_plot.png")
Results: